Antimicrobials design and development
The emergence of Multi-Drugs Resistant Bacteria is becoming a global emergency. The ability of bacteria to reduce their susceptibility to antimicrobial drugs importantly compromises the treatment of infectious diseases. Exposure to antibiotics induces the expression of mutagenic bacterial stress–response pathways, most importantly the SOS response.
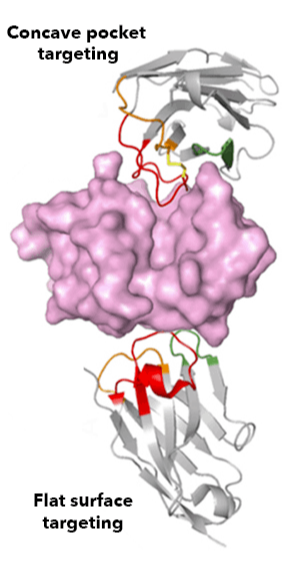
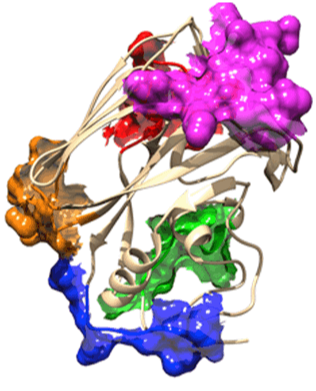
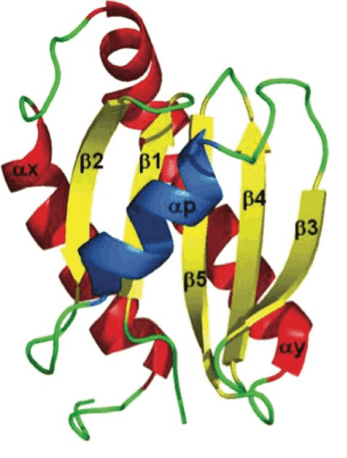
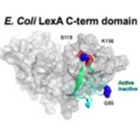
The aim of our study is to develop new suppressors of the SOS-machinery, through inhibition of LexA dissociation from DNA binding box, occurring upon autoproteolytic digestion of LexA, triggered by RecA recombinase. Both autoproteolysis inhibition and interference with RecA/LexA interaction could be explored [1, 2]. For our purposes, we will combine multiple approaches to obtain lead molecules active against SOS-response. Both in silico screenings and in vitro combinatorial methods, to generate and screen macrocycle libraries, are currently applied to our targets. Furthermore, the emerging powerful tool of camelids derived nanobodies has been explored with promising results. The capability of most promising leads (either synthetic compounds or nanobodies) to synergize with antibiotic treatments and keep SOS response repressed will be investigated in bacterial cultures models and by biochemical assays. The achievement of our goals will provide important clues in the development of novel strategies to suppress SOS pathway, rehabilitating in therapy dismissed antibiotics.